-Search query
-Search result
Showing 1 - 50 of 904 items for (author: smith & sm)

EMDB-42977: 
Cryo-EM structure of singly-bound SNF2h-nucleosome complex with SNF2h at inactive SHL2 (conformation 1)
Method: single particle / : Chio US, Palovcak E, Armache JP, Narlikar GJ, Cheng Y

EMDB-43000: 
Cryo-EM structure of SNF2h-nucleosome complex (consensus structure)
Method: single particle / : Chio US, Palovcak E, Armache JP, Narlikar GJ, Cheng Y

EMDB-43001: 
Cryo-EM structure of SNF2h-nucleosome complex (single-bound structure)
Method: single particle / : Chio US, Palovcak E, Armache JP, Narlikar GJ, Cheng Y

EMDB-43002: 
Cryo-EM structure of doubly-bound SNF2h-nucleosome complex
Method: single particle / : Chio US, Palovcak E, Armache JP, Narlikar GJ, Cheng Y

EMDB-43003: 
Cryo-EM structure of singly-bound SNF2h-nucleosome complex with SNF2h at inactive SHL2 (conformation 2)
Method: single particle / : Chio US, Palovcak E, Armache JP, Narlikar GJ, Cheng Y

EMDB-43004: 
Cryo-EM structure of doubly-bound SNF2h-nucleosome complex (conformation 1)
Method: single particle / : Chio US, Palovcak E, Armache JP, Narlikar GJ, Cheng Y

EMDB-43005: 
Cryo-EM structure of doubly-bound SNF2h-nucleosome complex (conformation 2)
Method: single particle / : Chio US, Palovcak E, Armache JP, Narlikar GJ, Cheng Y

PDB-8v4y: 
Cryo-EM structure of singly-bound SNF2h-nucleosome complex with SNF2h at inactive SHL2 (conformation 1)
Method: single particle / : Chio US, Palovcak E, Armache JP, Narlikar GJ, Cheng Y

PDB-8v6v: 
Cryo-EM structure of doubly-bound SNF2h-nucleosome complex
Method: single particle / : Chio US, Palovcak E, Armache JP, Narlikar GJ, Cheng Y

PDB-8v7l: 
Cryo-EM structure of singly-bound SNF2h-nucleosome complex with SNF2h at inactive SHL2 (conformation 2)
Method: single particle / : Chio US, Palovcak E, Armache JP, Narlikar GJ, Cheng Y

EMDB-41933: 
CryoEM map of horse spleen apoferritin determined as a reference for benchmarking square and rectangular apertures for cryo-EM
Method: single particle / : Brown HG, Hanssen E

EMDB-41936: 
CryoEM map of horse spleen apoferritin determined as a reference for benchmarking square and rectangular apertures for cryo-EM (Falcon IV round beam)
Method: single particle / : Brown HG, Hanssen E

EMDB-41937: 
CryoEM map of horse spleen apoferritin determined for benchmarking square and rectangular apertures for cryo-EM (Falcon IV square beam)
Method: single particle / : Brown HG, Hanssen E

EMDB-41938: 
CryoEM map of horse spleen apoferritin determined for benchmarking square and rectangular apertures for cryo-EM (Gatan K3 rectangular beam)
Method: single particle / : Brown HG, Hanssen E

EMDB-42882: 
Amylin 1 receptor bound to salmon calcitonin (Amy1R:sCT) reconstructed from cryo-EM datasets recorded using a rectangular aperture
Method: single particle / : Brown HG, Hanssen E
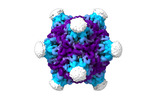
EMDB-19024: 
Structure of the PNMA2 capsid
Method: single particle / : Erlendsson S, Xu J, Shepherd JD, Briggs JAG
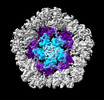
EMDB-19025: 
Structure of the five-fold capsomer of the PNMA2 capsid
Method: single particle / : Erlendsson S, Xu J, Shepherd JD, Briggs JAG
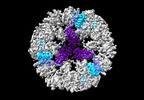
EMDB-19026: 
Structure of the three-fold capsomer of the PNMA2 capsid
Method: single particle / : Erlendsson S, Xu J, Shepherd JD, Briggs JAG
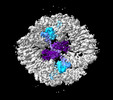
EMDB-19027: 
Structure of the two-fold capsomer of the PNMA2 capsid
Method: single particle / : Erlendsson S, Xu J, Shepherd JD, Briggs JAG

PDB-8rb3: 
Structure of the PNMA2 capsid
Method: single particle / : Erlendsson S, Xu J, Shepherd JD, Briggs JAG

PDB-8rb4: 
Structure of the five-fold capsomer of the PNMA2 capsid
Method: single particle / : Erlendsson S, Xu J, Shepherd JD, Briggs JAG

PDB-8rb5: 
Structure of the three-fold capsomer of the PNMA2 capsid
Method: single particle / : Erlendsson S, Xu J, Shepherd JD, Briggs JAG

PDB-8rb7: 
Structure of the two-fold capsomer of the PNMA2 capsid
Method: single particle / : Erlendsson S, Xu J, Shepherd JD, Briggs JAG

EMDB-19194: 
In situ cryo-electron tomogram of an autophagosome in the projection of an iPSC-derived neuron #2
Method: electron tomography / : Hoyer MJ, Capitanio C, Smith IR, Paoli JC, Bieber A, Jiang Y, Paulo JA, Gonzalez-Lozano MA, Baumeister W, Wilfling F, Schulman BA, Harper WJ

EMDB-19346: 
In situ cryo-electron tomogram of an autophagosome in the projection of an iPSC-derived neuron #1
Method: electron tomography / : Hoyer MJ, Capitanio C, Smith IR, Paoli JC, Bieber A, Jiang Y, Paulo JA, Gonzalez-Lozano MA, Baumeister W, Wilfling F, Schulman BA, Harper WJ

EMDB-42600: 
Murine norovirus in the presence of 1mM calcium
Method: single particle / : Smith TJ

EMDB-42604: 
Murine norovirus + 1 mM MgCl2
Method: single particle / : Smith TJ

EMDB-42623: 
Murine norovirus dialyzed against EDTA
Method: single particle / : Smith TJ

PDB-8uux: 
Murine norovirus capsid protein in the presence of 1mM calcium
Method: single particle / : Smith TJ

EMDB-37179: 
Post-processed counting mode Adeno-associated virus (AAV) (RELION)
Method: single particle / : Burton-Smith RN, Murata K

EMDB-37180: 
Post-processed PASR pre-processed Adeno-associated virus (AAV) (RELION)
Method: single particle / : Burton-Smith RN, Murata K

EMDB-37175: 
Apoferritin (8K super resolution from 32 Falcon 4 micrographs)
Method: single particle / : Burton-Smith RN, Murata K

EMDB-37173: 
Reprocessing of Fourier binned movies of EMPIAR-10216 Apoferritin for PASR validation tests
Method: single particle / : Burton-Smith RN, Murata K

EMDB-37169: 
Melbournevirus (whole virus, Bayesian polished reconstruction, realigned to I1 symmetry)
Method: single particle / : Burton-Smith RN, Murata K

EMDB-37181: 
Nodavirus (reprocessed EMPIAR-10203) for single-frame PASR comparison
Method: single particle / : Burton-Smith RN, Murata K

EMDB-37186: 
Jackbean Urease (Reprocessing of EMPIAR-10549 for comparison to PASR processing)
Method: single particle / : Burton-Smith RN, Murata K

EMDB-37174: 
PASR pre-processed Apoferritin (PASR 4K counting mode from 32 Falcon 4 micrographs)
Method: single particle / : Burton-Smith RN, Murata K

EMDB-37190: 
Five-fold Melbournevirus block (PASR pre-processed K2 counting mode data)
Method: single particle / : Burton-Smith RN, Murata K

EMDB-37189: 
Three-fold Melbournevirus block (PASR pre-processed K2 counting mode data)
Method: single particle / : Burton-Smith RN, Murata K

EMDB-37172: 
Reprocessing of EMPIAR-10216 Apoferritin for PASR validation tests
Method: single particle / : Burton-Smith RN, Murata K

EMDB-37177: 
Apoferritin (8K super resolution from 32 Falcon 4 micrographs) processed with CryoSPARC
Method: single particle / : Burton-Smith RN, Murata K

EMDB-37183: 
Nodavirus (Fourier cropped EMPIAR-10203) for single-frame PASR comparison
Method: single particle / : Burton-Smith RN, Murata K

EMDB-37185: 
Jackbean Urease (Fourier cropped K3 super resolution then PASR pre-processed) (Original data: EMPIAR-10549)
Method: single particle / : Burton-Smith RN, Murata K

EMDB-37176: 
Apoferritin (4K counting mode from 32 Falcon 4 micrographs)
Method: single particle / : Burton-Smith RN, Murata K

EMDB-37187: 
Jackbean Urease (Fourier cropped K3 super resolution movies) (Based on EMPIAR-10549)
Method: single particle / : Burton-Smith RN, Murata K

EMDB-37182: 
Nodavirus (Fourier cropped then PASR pre-processed EMPIAR-10203) for single-frame PASR comparison
Method: single particle / : Burton-Smith RN, Murata K

EMDB-37171: 
PASR (Post-acquisition Super Resolution) reprocessing of EMPIAR-10216 Apoferritin
Method: single particle / : Burton-Smith RN, Murata K

EMDB-37178: 
PASR pre-processed Apoferritin (PASR 4K counting mode from 32 Falcon 4 micrographs) with CryoSPARC
Method: single particle / : Burton-Smith RN, Murata K

EMDB-37188: 
Two-fold Melbournevirus block (Reprocessing via PASR pre-processed K2 counting mode data)
Method: single particle / : Burton-Smith RN, Murata K
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search




 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
